|
| 1 | +# --- |
| 2 | +# jupyter: |
| 3 | +# jupytext: |
| 4 | +# formats: py:percent,ipynb |
| 5 | +# text_representation: |
| 6 | +# extension: .py |
| 7 | +# format_name: percent |
| 8 | +# format_version: '1.3' |
| 9 | +# jupytext_version: 1.14.5 |
| 10 | +# kernelspec: |
| 11 | +# display_name: Python 3 (ipykernel) |
| 12 | +# language: python |
| 13 | +# name: python3 |
| 14 | +# --- |
| 15 | + |
| 16 | +# %% |
| 17 | +# Copyright (c) MONAI Consortium |
| 18 | +# Licensed under the Apache License, Version 2.0 (the "License"); |
| 19 | +# you may not use this file except in compliance with the License. |
| 20 | +# You may obtain a copy of the License at |
| 21 | +# http://www.apache.org/licenses/LICENSE-2.0 |
| 22 | +# Unless required by applicable law or agreed to in writing, software |
| 23 | +# distributed under the License is distributed on an "AS IS" BASIS, |
| 24 | +# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. |
| 25 | +# See the License for the specific language governing permissions and |
| 26 | +# limitations under the License. |
| 27 | + |
| 28 | +# %% [markdown] |
| 29 | +# # Diffusion Autoencoder Tutorial with Image Manipulation |
| 30 | +# |
| 31 | +# This tutorial illustrates how to use MONAI Generative Models for training a 2D Diffusion Autoencoder model [1]. |
| 32 | +# |
| 33 | +# In summary, the tutorial will cover the following: |
| 34 | +# 1. Loading and preprocessing a dataset (we extract the brain MRI dataset 2D slices from 3D volumes from the BraTS dataset); |
| 35 | +# 2. Training a 2D diffusion model and semantic encoder with a ResNet18 backbone; |
| 36 | +# 3. Evaluate the learned latent space by applying a Logistic Regression classifier and manipulating images with using the direction learned by the classifier. |
| 37 | +# |
| 38 | +# |
| 39 | +# During inference, the model encodes the image to latent space which is used as conditioning for the diffusion unet diffusion model. [1] trains a latent diffusion model for being able to generate unconditional images, which we do not do in this tutorial. Here, we are interested in checking wether the latent space contains classification information or not and if can be used to manipulate the images. |
| 40 | +# |
| 41 | +# [1] Preechakul et al. [Diffusion Autoencoders: Toward a Meaningful and Decodable Representation](https://arxiv.org/abs/2111.15640). CVPR 2022 |
| 42 | +# |
| 43 | +# 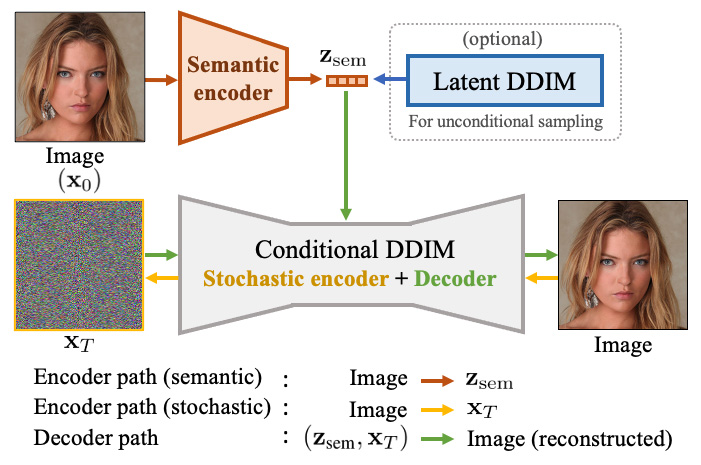 |
| 44 | + |
| 45 | +# %% [markdown] |
| 46 | +# ## Setup imports |
| 47 | + |
| 48 | +# %% jupyter={"outputs_hidden": false} |
| 49 | +import os |
| 50 | +import tempfile |
| 51 | +import time |
| 52 | +import os |
| 53 | +import matplotlib.pyplot as plt |
| 54 | +import numpy as np |
| 55 | +import torch |
| 56 | +import torch.nn.functional as F |
| 57 | +import torchvision |
| 58 | +import sys |
| 59 | +from monai import transforms |
| 60 | +from monai.apps import DecathlonDataset |
| 61 | +from monai.config import print_config |
| 62 | +from monai.data import DataLoader |
| 63 | +from monai.utils import set_determinism |
| 64 | +from torch.cuda.amp import GradScaler, autocast |
| 65 | +from tqdm import tqdm |
| 66 | +from sklearn.linear_model import LogisticRegression |
| 67 | + |
| 68 | +from generative.inferers import DiffusionInferer |
| 69 | +from generative.networks.nets.diffusion_model_unet import DiffusionModelUNet |
| 70 | +from generative.networks.schedulers.ddim import DDIMScheduler |
| 71 | + |
| 72 | +torch.multiprocessing.set_sharing_strategy("file_system") |
| 73 | + |
| 74 | +print_config() |
| 75 | + |
| 76 | + |
| 77 | +# %% [markdown] |
| 78 | +# ## Setup data directory |
| 79 | + |
| 80 | +# %% jupyter={"outputs_hidden": false} |
| 81 | +directory = os.environ.get("MONAI_DATA_DIRECTORY") |
| 82 | +root_dir = tempfile.mkdtemp() if directory is None else directory |
| 83 | +root_dir = '/home/s2086085/pedro_idcom/experiment_data' |
| 84 | + |
| 85 | +# %% [markdown] |
| 86 | +# ## Set deterministic training for reproducibility |
| 87 | + |
| 88 | +# %% jupyter={"outputs_hidden": false} |
| 89 | +set_determinism(42) |
| 90 | + |
| 91 | +# %% [markdown] |
| 92 | +# # Dataset preparation |
| 93 | +# |
| 94 | +# ## Setup BRATS Dataset - Transforms for extracting 2D slices from 3D volumes |
| 95 | +# |
| 96 | +# We now download the BraTS dataset and extract the 2D slices from the 3D volumes. The `slice_label` is used to indicate whether the slice contains an anomaly or not. |
| 97 | + |
| 98 | +# %% [markdown] |
| 99 | +# Here we use transforms to augment the training dataset, as usual: |
| 100 | +# |
| 101 | +# 1. `LoadImaged` loads the brain images from files. |
| 102 | +# 2. `EnsureChannelFirstd` ensures the original data to construct "channel first" shape. |
| 103 | +# 3. The first `Lambdad` transform chooses the first channel of the image, which is the T1-weighted image. |
| 104 | +# 4. `Spacingd` resamples the image to the specified voxel spacing, we use 3,3,2 mm to match the original paper. |
| 105 | +# 5. `ScaleIntensityRangePercentilesd` Apply range scaling to a numpy array based on the intensity distribution of the input. Transform is very common with MRI images. |
| 106 | +# 6. `RandSpatialCropd` randomly crop out a 2D patch from the 3D image. |
| 107 | +# 6. The last `Lambdad` transform obtains `slice_label` by summing up the label to have a single scalar value (healthy `=1` or not `=2` ). |
| 108 | + |
| 109 | +# %% |
| 110 | +channel = 0 # 0 = Flair |
| 111 | +assert channel in [0, 1, 2, 3], "Choose a valid channel" |
| 112 | + |
| 113 | +train_transforms = transforms.Compose( |
| 114 | + [ |
| 115 | + transforms.LoadImaged(keys=["image", "label"]), |
| 116 | + transforms.EnsureChannelFirstd(keys=["image", "label"]), |
| 117 | + transforms.Lambdad(keys=["image"], func=lambda x: x[channel, :, :, :]), |
| 118 | + transforms.AddChanneld(keys=["image"]), |
| 119 | + transforms.EnsureTyped(keys=["image", "label"]), |
| 120 | + transforms.Orientationd(keys=["image", "label"], axcodes="RAS"), |
| 121 | + transforms.Spacingd(keys=["image", "label"], pixdim=(3.0, 3.0, 2.0), mode=("bilinear", "nearest")), |
| 122 | + transforms.CenterSpatialCropd(keys=["image", "label"], roi_size=(64, 64, 44)), |
| 123 | + transforms.ScaleIntensityRangePercentilesd(keys="image", lower=0, upper=99.5, b_min=0, b_max=1), |
| 124 | + transforms.RandSpatialCropd(keys=["image", "label"], roi_size=(64, 64, 1), random_size=False), |
| 125 | + transforms.Lambdad(keys=["image", "label"], func=lambda x: x.squeeze(-1)), |
| 126 | + transforms.CopyItemsd(keys=["label"], times=1, names=["slice_label"]), |
| 127 | + transforms.Lambdad(keys=["slice_label"], func=lambda x: 1.0 if x.sum() > 0 else 0.0), |
| 128 | + ] |
| 129 | +) |
| 130 | + |
| 131 | +# %% [markdown] |
| 132 | +# ### Load Training and Validation Datasets |
| 133 | + |
| 134 | +# %% jupyter={"outputs_hidden": false} |
| 135 | +train_ds = DecathlonDataset( |
| 136 | + root_dir=root_dir, |
| 137 | + task="Task01_BrainTumour", |
| 138 | + section="training", |
| 139 | + cache_rate=1.0, # you may need a few Gb of RAM... Set to 0 otherwise |
| 140 | + num_workers=4, |
| 141 | + download=False, # Set download to True if the dataset hasnt been downloaded yet |
| 142 | + seed=0, |
| 143 | + transform=train_transforms, |
| 144 | +) |
| 145 | +print(f"Length of training data: {len(train_ds)}") |
| 146 | +print(f'Train image shape {train_ds[0]["image"].shape}') |
| 147 | + |
| 148 | +# %% |
| 149 | +val_ds = DecathlonDataset( |
| 150 | + root_dir=root_dir, |
| 151 | + task="Task01_BrainTumour", |
| 152 | + section="validation", |
| 153 | + cache_rate=1, # you may need a few Gb of RAM... Set to 0 otherwise |
| 154 | + num_workers=4, |
| 155 | + download=False, # Set download to True if the dataset hasnt been downloaded yet |
| 156 | + seed=0, |
| 157 | + transform=train_transforms, |
| 158 | +) |
| 159 | +print(f"Length of training data: {len(val_ds)}") |
| 160 | +print(f'Validation Image shape {val_ds[0]["image"].shape}') |
| 161 | + |
| 162 | + |
| 163 | +# %% [markdown] |
| 164 | +# # Training |
| 165 | +# ## Define network, scheduler, optimizer, and inferer |
| 166 | +# |
| 167 | +# At this step, we instantiate the MONAI components to create a DDIM, the UNET with conditioning, the noise scheduler, and the inferer used for training and sampling. We are using |
| 168 | +# the deterministic DDIM scheduler containing 1000 timesteps, and a 2D UNET with attention mechanisms. |
| 169 | +# |
| 170 | +# The `attention` mechanism is essential for ensuring good conditioning and images manipulation here. |
| 171 | +# |
| 172 | +# The `embedding_dimension` parameter controls the dimension of the latent dimension learned by the semantic encoder. |
| 173 | +# |
| 174 | + |
| 175 | +# %% jupyter={"outputs_hidden": false} |
| 176 | +class Diffusion_AE(torch.nn.Module): |
| 177 | + def __init__(self, embedding_dimension = 64): |
| 178 | + super().__init__() |
| 179 | + self.unet = DiffusionModelUNet( |
| 180 | + spatial_dims=2, |
| 181 | + in_channels=1, |
| 182 | + out_channels=1, |
| 183 | + num_channels=(128, 256, 256), |
| 184 | + attention_levels=(False, True, True), |
| 185 | + num_res_blocks=1, |
| 186 | + num_head_channels=64, |
| 187 | + with_conditioning=True, |
| 188 | + cross_attention_dim=1, |
| 189 | + ) |
| 190 | + self.semantic_encoder = torchvision.models.resnet18() |
| 191 | + self.semantic_encoder.conv1 = torch.nn.Conv2d(1, 64, kernel_size=7, stride=2, padding=3, bias=False) |
| 192 | + self.semantic_encoder.fc = torch.nn.Linear(512, embedding_dimension) |
| 193 | + |
| 194 | + |
| 195 | + def forward(self, xt, x_cond, t): |
| 196 | + latent = self.semantic_encoder(x_cond) |
| 197 | + noise_pred = self.unet(x=xt, timesteps=t, context=latent.unsqueeze(2)) |
| 198 | + return noise_pred, latent |
| 199 | + |
| 200 | +device = torch.device("cuda:2") |
| 201 | +model = Diffusion_AE(embedding_dimension = 512).to(device) |
| 202 | +scheduler = DDIMScheduler(num_train_timesteps=1000) |
| 203 | +optimizer = torch.optim.Adam(params=model.parameters(), lr=1e-5) |
| 204 | +inferer = DiffusionInferer(scheduler) |
| 205 | + |
| 206 | + |
| 207 | +# %% [markdown] |
| 208 | +# ## Training a diffusion model and semantic encoder |
| 209 | + |
| 210 | +# %% |
| 211 | +n_iterations = 1e4 # training for longer helps a lot with reconstruction quality, even if the loss is already low |
| 212 | +batch_size = 64 |
| 213 | +val_interval = 100 |
| 214 | +iter_loss_list, val_iter_loss_list = [], [] |
| 215 | +iterations = [] |
| 216 | +iteration, iter_loss = 0, 0 |
| 217 | + |
| 218 | +train_loader = DataLoader( |
| 219 | + train_ds, batch_size=batch_size, shuffle=True, num_workers=4, drop_last=True, persistent_workers=True |
| 220 | +) |
| 221 | +val_loader = DataLoader( |
| 222 | + val_ds, batch_size=batch_size, shuffle=False, num_workers=4, drop_last=True, persistent_workers=True |
| 223 | +) |
| 224 | + |
| 225 | +total_start = time.time() |
| 226 | + |
| 227 | +while iteration < n_iterations: |
| 228 | + for batch in train_loader: |
| 229 | + iteration += 1 |
| 230 | + model.train() |
| 231 | + optimizer.zero_grad(set_to_none=True) |
| 232 | + images= batch["image"].to(device) |
| 233 | + noise = torch.randn_like(images).to(device) |
| 234 | + # Create timesteps |
| 235 | + timesteps = torch.randint(0, inferer.scheduler.num_train_timesteps, (batch_size,)).to(device).long() |
| 236 | + # Get model prediction |
| 237 | + # cross attention expects shape [batch size, sequence length, channels], we are use channels = latent dimension and sequence length = 1 |
| 238 | + latent = model.semantic_encoder(images) |
| 239 | + noise_pred = inferer(inputs=images, diffusion_model=model.unet, noise=noise, timesteps=timesteps, condition = latent.unsqueeze(2)) |
| 240 | + loss = F.mse_loss(noise_pred.float(), noise.float()) |
| 241 | + |
| 242 | + loss.backward() |
| 243 | + optimizer.step() |
| 244 | + |
| 245 | + iter_loss += loss.item() |
| 246 | + sys.stdout.write(f"Iteration {iteration}/{n_iterations} - train Loss {loss.item():.4f}" + "\r") |
| 247 | + sys.stdout.flush() |
| 248 | + |
| 249 | + if (iteration) % val_interval == 0: |
| 250 | + model.eval() |
| 251 | + val_iter_loss = 0 |
| 252 | + for val_step, val_batch in enumerate(val_loader): |
| 253 | + with torch.no_grad(): |
| 254 | + images = val_batch["image"].to(device) |
| 255 | + timesteps = torch.randint(0, inferer.scheduler.num_train_timesteps, (batch_size,)).to(device).long() |
| 256 | + noise = torch.randn_like(images).to(device) |
| 257 | + latent = model.semantic_encoder(images) |
| 258 | + noise_pred = inferer(inputs=images, diffusion_model=model.unet, noise=noise, timesteps=timesteps, condition = latent.unsqueeze(2)) |
| 259 | + val_loss = F.mse_loss(noise_pred.float(), noise.float()) |
| 260 | + |
| 261 | + val_iter_loss += val_loss.item() |
| 262 | + iter_loss_list.append(iter_loss / val_interval) |
| 263 | + val_iter_loss_list.append(val_iter_loss / (val_step + 1)) |
| 264 | + iterations.append(iteration) |
| 265 | + iter_loss = 0 |
| 266 | + print(f"Iteration {iteration} - Interval Loss {iter_loss_list[-1]:.4f}, Interval Loss Val {val_iter_loss_list[-1]:.4f}") |
| 267 | + |
| 268 | + |
| 269 | +total_time = time.time() - total_start |
| 270 | + |
| 271 | +print(f"train diffusion completed, total time: {total_time}.") |
| 272 | + |
| 273 | +plt.style.use("seaborn-bright") |
| 274 | +plt.title("Learning Curves Diffusion Model", fontsize=20) |
| 275 | + |
| 276 | +plt.plot(iterations, iter_loss_list, color="C0", linewidth=2.0, label="Train") |
| 277 | +plt.plot(iterations, val_iter_loss_list, color="C4", linewidth=2.0, label="Validation") |
| 278 | + |
| 279 | +plt.yticks(fontsize=12), plt.xticks(fontsize=12) |
| 280 | +plt.xlabel("Iterations", fontsize=16), plt.ylabel("Loss", fontsize=16) |
| 281 | +plt.legend(prop={"size": 14}) |
| 282 | +plt.show() |
| 283 | + |
| 284 | +# %% [markdown] |
| 285 | +# # Evaluation |
| 286 | +# ## Generate synthetic data |
| 287 | +# |
| 288 | +# We use the semantic encoder to get a latent space from the validation dataset. We then use the diffusion model to generate synthetic data from the latent space. |
| 289 | + |
| 290 | +# %% |
| 291 | +scheduler.set_timesteps(num_inference_steps=100) |
| 292 | +batch = next(iter(val_loader)) |
| 293 | +images = batch["image"].to(device) |
| 294 | +noise = torch.randn_like(images).to(device) |
| 295 | +latent = model.semantic_encoder(images) |
| 296 | +reconstruction = inferer.sample(input_noise=noise, diffusion_model=model.unet, scheduler=scheduler, save_intermediates=False, conditioning=latent.unsqueeze(2)) |
| 297 | + |
| 298 | +# %% |
| 299 | +grid = torchvision.utils.make_grid(torch.cat([images[:8],reconstruction[:8]]), nrow=8, padding=2, normalize=True, range=None, scale_each=False, pad_value=0) |
| 300 | +plt.figure(figsize=(15,5)) |
| 301 | +plt.imshow(grid.detach().cpu().numpy()[0], cmap= 'gray') |
| 302 | +plt.axis('off') |
| 303 | + |
| 304 | +# %% [markdown] |
| 305 | +# ## Evaluate Latent Space |
| 306 | +# |
| 307 | +# First, obtain the latent space from the entire training and validation datasets. |
| 308 | +# Then, we train a logistic regression classifier on the latent space. |
| 309 | + |
| 310 | +# %% |
| 311 | +# get latent space of training set |
| 312 | +latents_train = [] |
| 313 | +classes_train = [] |
| 314 | +for i in range(15): # 15 slices from each volume |
| 315 | + for batch in train_loader: |
| 316 | + images = batch["image"].to(device) |
| 317 | + latent = model.semantic_encoder(images) |
| 318 | + latents_train.append(latent.detach().cpu().numpy()) |
| 319 | + classes_train.append(batch["slice_label"].numpy()) |
| 320 | + |
| 321 | +latents_train = np.concatenate(latents_train, axis=0) |
| 322 | +classes_train = np.concatenate(classes_train, axis=0) |
| 323 | + |
| 324 | +# get latent space of validation set |
| 325 | +latents_val = [] |
| 326 | +classes_val = [] |
| 327 | +for batch in val_loader: |
| 328 | + images = batch["image"].to(device) |
| 329 | + latent = model.semantic_encoder(images) |
| 330 | + latents_val.append(latent.detach().cpu().numpy()) |
| 331 | + classes_val.append(batch["slice_label"].numpy()) |
| 332 | +latents_val = np.concatenate(latents_val, axis=0) |
| 333 | +classes_val = np.concatenate(classes_val, axis=0) |
| 334 | + |
| 335 | +# %% |
| 336 | +latents_train.shape , classes_train.shape |
| 337 | + |
| 338 | +# %% |
| 339 | +clf = LogisticRegression(solver = 'newton-cg', random_state=0).fit(latents_train, classes_train) |
| 340 | +clf.score(latents_train, classes_train), clf.score(latents_val, classes_val) |
| 341 | + |
| 342 | +# %% |
| 343 | +w = torch.Tensor(clf.coef_).float().to(device) |
| 344 | + |
| 345 | +# %% [markdown] |
| 346 | +# ## Manipulate Latent Space |
| 347 | +# |
| 348 | +# The logistic regression classifier learns a direction `w` in the latent space that separates the healthy and abnormal slices. We can use this direction to manipulate the images by `latent = latent + s * w`, where `s` controls the strength of the manipulation. Negative `s` will move the image in the opposite direction. |
| 349 | + |
| 350 | +# %% |
| 351 | +s = -1.5 |
| 352 | + |
| 353 | +scheduler.set_timesteps(num_inference_steps=100) |
| 354 | +batch = next(iter(val_loader)) |
| 355 | +images = batch["image"].to(device) |
| 356 | +noise = torch.randn_like(images).to(device) |
| 357 | + |
| 358 | +latent = model.semantic_encoder(images) |
| 359 | + |
| 360 | +latent_manip = latent + s * w[0] |
| 361 | + |
| 362 | +reconstruction = inferer.sample(input_noise=noise, diffusion_model=model.unet, scheduler=scheduler, save_intermediates=False, conditioning=latent.unsqueeze(1)) |
| 363 | +manipulated_images = inferer.sample(input_noise=noise, diffusion_model=model.unet, scheduler=scheduler, save_intermediates=False, conditioning=latent_manip.unsqueeze(1)) |
| 364 | + |
| 365 | +# %% |
| 366 | +nb = 8 |
| 367 | +grid = torchvision.utils.make_grid(torch.cat([images[:nb], reconstruction[:nb], manipulated_images[:nb]]), |
| 368 | + nrow=8, normalize=False, range=None, scale_each=False, pad_value=0) |
| 369 | +plt.figure(figsize=(15,5)) |
| 370 | +plt.imshow(grid.detach().cpu().numpy()[0], cmap= 'gray') |
| 371 | +plt.axis('off') |
| 372 | +plt.title(f"Original, Reconstruction, Manipulated s = {s}"); |
0 commit comments